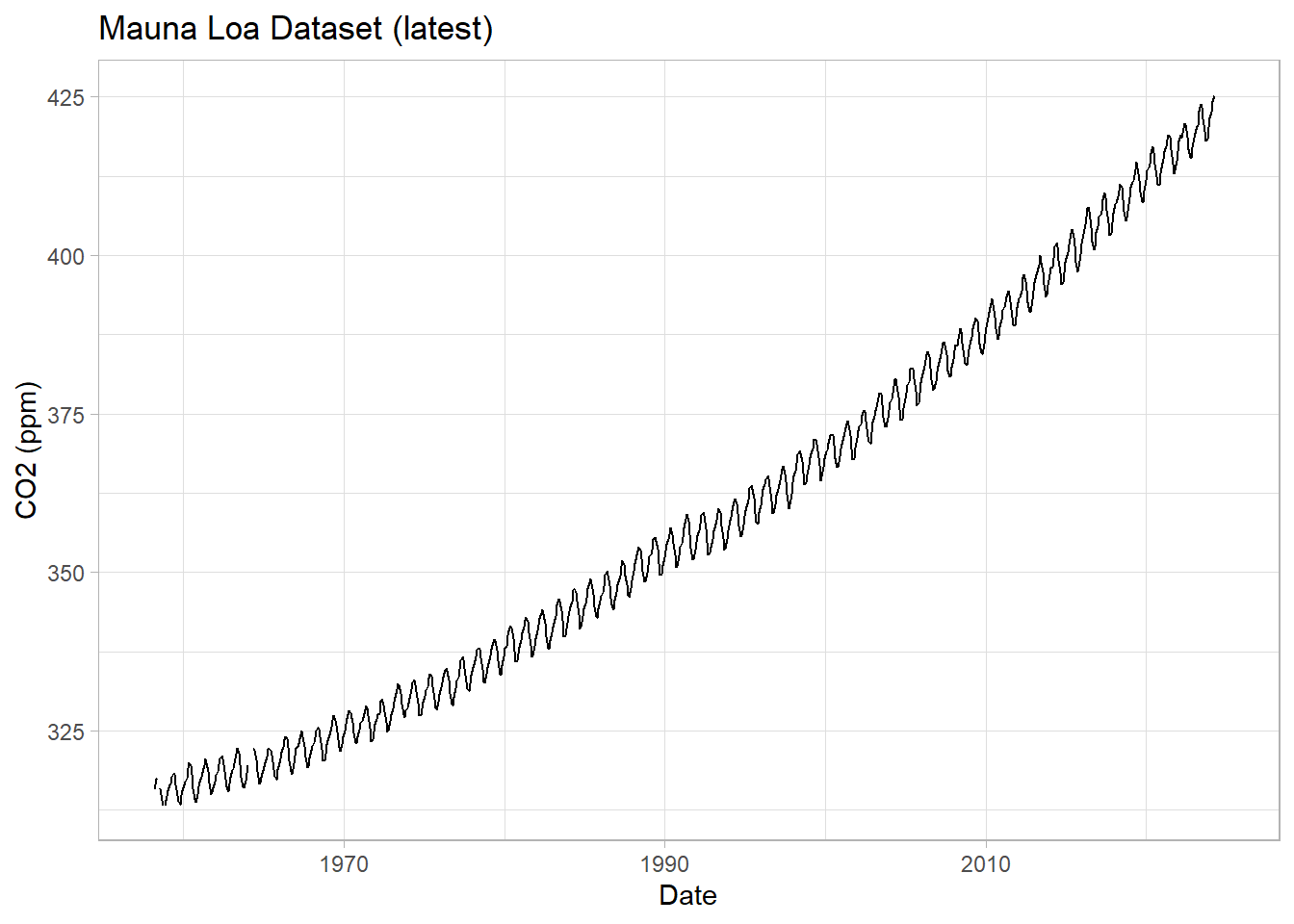2.2 Data Wrangling
2.2.2 Load data
We can pull the data directly from the Scripps Institute website
df1 <- read.csv("https://scrippsco2.ucsd.edu/assets/data/atmospheric/stations/in_situ_co2/monthly/monthly_in_situ_co2_mlo.csv")
head(df1)## X...........................................................................................
## 1 Atmospheric CO2 concentrations (ppm) derived from in situ air measurements
## 2 at Mauna Loa, Observatory, Hawaii: Latitude 19.5°N Longitude 155.6°W Elevation 3397m
## 3 Since December 2022 sampling has temporarily been relocated to MaunuaKea, Hawaii
## 4 Latitude 19.8°N Longitude 155.5°W Elevation 4145m
## 5
## 6 Source: R. F. Keeling, S. J. Walker, S. C. Piper and A. F. BollenbacherOoo, that doesn’t look right. This file has a big header, and doesn’t quite follow a normal spreadsheet format for the column headers. Let’s skip the header and column names. We can add them back in after loading.
df1 <- read.csv("https://scrippsco2.ucsd.edu/assets/data/atmospheric/stations/in_situ_co2/monthly/monthly_in_situ_co2_mlo.csv",skip=64,header = F)
head(df1)## V1 V2 V3 V4 V5 V6 V7 V8 V9 V10 V11
## 1 1958 1 21200 1958.041 -99.99 -99.99 -99.99 -99.99 -99.99 -99.99 MLO
## 2 1958 2 21231 1958.126 -99.99 -99.99 -99.99 -99.99 -99.99 -99.99 MLO
## 3 1958 3 21259 1958.203 315.71 314.44 316.20 314.91 315.71 314.44 MLO
## 4 1958 4 21290 1958.288 317.45 315.16 317.30 314.99 317.45 315.16 MLO
## 5 1958 5 21320 1958.370 317.51 314.69 317.89 315.07 317.51 314.69 MLO
## 6 1958 6 21351 1958.455 -99.99 -99.99 317.27 315.15 317.27 315.15 MLOOkay, not the data looks right.
2.2.3 Format data
Lets add the column names.
names(df1) <- c('Yr','Mn','XLDate','Date','CO2','seasonally adjusted','fit','seasonally adjusted fit','CO2 filled','seasonally adjusted filled','Sta')
head(df1)## Yr Mn XLDate Date CO2 seasonally adjusted fit seasonally adjusted fit
## 1 1958 1 21200 1958.041 -99.99 -99.99 -99.99 -99.99
## 2 1958 2 21231 1958.126 -99.99 -99.99 -99.99 -99.99
## 3 1958 3 21259 1958.203 315.71 314.44 316.20 314.91
## 4 1958 4 21290 1958.288 317.45 315.16 317.30 314.99
## 5 1958 5 21320 1958.370 317.51 314.69 317.89 315.07
## 6 1958 6 21351 1958.455 -99.99 -99.99 317.27 315.15
## CO2 filled seasonally adjusted filled Sta
## 1 -99.99 -99.99 MLO
## 2 -99.99 -99.99 MLO
## 3 315.71 314.44 MLO
## 4 317.45 315.16 MLO
## 5 317.51 314.69 MLO
## 6 317.27 315.15 MLOThe missing value default here is -99.99, let’s change that to NA, R’s standard.